Swot L2_LR_SSH#
This chapter will present the functionalities specific to the Level 2 SWOT Low Rate products.
from fcollections.implementations import (
# Handler
NetcdfFilesDatabaseSwotLRL2,
# Version
L2Version, Timeliness)
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import cartopy.crs as ccrs
Data samples#
We will illustrate the functionalities using a data sample from AVISO. You can use the altimetry-downloader-aviso tool to run the following script.
import logging
from pathlib import Path
from altimetry_downloader_aviso import get
logging.basicConfig()
logging.getLogger("altimetry_downloader_aviso").setLevel("INFO")
DATA_DIR = Path(__file__).resolve().parent.parent / "data_l2_lr_ssh"
DATA_DIR.mkdir(exist_ok=True)
if __name__ == "__main__":
get(
"SWOT_L2_LR_SSH_Basic",
output_dir=DATA_DIR,
cycle_number=[9, 10, 11],
pass_number=[10, 11],
version="P?C?",
)
get(
"SWOT_L2_LR_SSH_Unsmoothed",
output_dir=DATA_DIR,
cycle_number=9,
pass_number=10,
version="PGC?",
)
Query overview#
Detailed information on the filters and reading arguments can be found in the
query API description
fcollections.implementations.NetcdfFilesDatabaseSwotLRL2.query()
The following examples can be used to build complex queries
A unique half orbit
fc.query(cycle_number=1, pass_number=1)
One half orbit repeating over all cycles
fc.query(pass_number=1)
A list of half orbits, over multiple cycles
fc.query(cycle_number=slice(1, 4), pass_number=[1, 3])
A specific orbit of the SWOT mission
fc.query(phase='CALVAL')
A time stamp
fc.query(time='2024-01-01')
A period
fc.query(time=('2024-01-01', '2024-03-31'))
A subset of variables
fc.query(selected_variables=['time', 'longitude', 'latitude'])
Note
Available variables can explored using
fcollections.implementations.NetcdfFilesDatabaseSwotLRL2.variables_info()
Zoom over an area selection
fc.query(bbox=(-10, 5, 35, 40))
Left swath (Unsmoothed only)
fc.query(left_swath=True, right_swath=False)
Right swath (Unsmoothed only)
fc.query(left_swath=False, right_swath=True)
Stacking over cycles (Basic, Expert, Windwave only)
fc.query(stack='CYCLES')
Stacking over both cycles and passes (Basic, Expert, Windwave only)
fc.query(stack='CYCLES_PASSES')
Use baseline C versions
fc.query(version='P?C?')
Use a specific baseline, and only reprocessed data
fc.query(version='PGD?')
Complete version specification
fc.query(version='PGD0_02')
Choose one dataset
fc.query(subset='Expert')
Generic information#
Generic information about the files set can be extracted. This available
information is specific to the mixins used to build
fcollections.implementations.NetcdfFilesDatabaseSwotLRL2
Time coverage
fc.time_coverage(subset='Expert', version='P?D?', phase='SCIENCE')
Time holes
fc.time_holes(subset='Expert', version='P?D?', phase='SCIENCE')
Half orbit range
fc.half_orbit_range(subset='Expert', version='P?D?', phase='SCIENCE')
Cycle range
fc.cycle_range(subset='Expert', version='P?D?', phase='SCIENCE')
Stack for temporal analysis#
The most prominent functionality is the ability to stack the half orbits when
the grid is fixed (Basic, Expert and WindWave subsets). This allows
to work along the cycle_number dimension and compute temporal analysis
(mean, standard deviation, …).
There are currently three modes for stacking the half orbits
NOSTACK: do not stack the half orbitsCYCLES: concatenate the half orbits of one cycle along thenum_linesdimension, and stack the cycles along a newcycle_numberdimensionCYCLES_PASSES: stack the half orbits along thecycle_numberandpass_numberdimensions. Useful for regional analysis where the half orbits are cropped and we need an additional dimension to reflect the spatial jump
fc = NetcdfFilesDatabaseSwotLRL2("data_l2_lr_ssh")
ds = fc.query(stack='CYCLES', cycle_number=[9, 10, 11], pass_number=10, subset='Basic')
ds.ssha_karin_2.data
|
||||||||||||||||
ds = fc.query(stack='CYCLES_PASSES', cycle_number=[9, 10, 11], pass_number=[10, 11], subset='Basic')
ds.ssha_karin_2.data
|
||||||||||||||||
Note
Incomplete cycles are completed with invalids
Filter Level-2 version#
The Level-2 version is a complex tag composed of a temporality (forward I or
reprocessed G), a baseline (major version A, B, C, D, …), a minor version
(0, 1, 2, ..) and a product counter (01, 02, …). The
fcollections.implementations.L2Version class can handle the tag
information and filter out non-desired versions. It can be partially initialized
in order to control the granularity of the filter.
version = L2Version(temporality=Timeliness.I)
version
PI??
fc.list_files(cycle_number=10, pass_number=10, version=version)
| cycle_number | pass_number | time | subset | version | filename | |
|---|---|---|---|---|---|---|
| 0 | 10 | 10 | [2024-01-25T08:02:33.000000, 2024-01-25T08:54:... | ProductSubset.Basic | PIC0_01 | /home/runner/work/fcollections/fcollections/do... |
version = L2Version(baseline='C')
version
P?C?
fc.list_files(cycle_number=9, pass_number=10, version=version)
| cycle_number | pass_number | time | subset | version | filename | |
|---|---|---|---|---|---|---|
| 0 | 9 | 10 | [2024-01-04T11:17:28.000000, 2024-01-04T12:08:... | ProductSubset.Unsmoothed | PGC0_01 | /home/runner/work/fcollections/fcollections/do... |
| 1 | 9 | 10 | [2024-01-04T11:17:27.000000, 2024-01-04T12:08:... | ProductSubset.Basic | PGC0_01 | /home/runner/work/fcollections/fcollections/do... |
| 2 | 9 | 10 | [2024-01-04T11:17:27.000000, 2024-01-04T12:08:... | ProductSubset.Basic | PIC0_01 | /home/runner/work/fcollections/fcollections/do... |
Area selection#
It is possible to select data crossing a specific region by providing bbox parameter to query or list_files method.
The bounding box is represented by a tuple of 4 float numbers, such as : (longitude_min, latitude_min, longitude_max, latitude_max). Its longitude must follow one of the known conventions: [0, 360[ or [-180, 180[.
If bbox’s longitude crosses -180/180, data around the crossing and matching the bbox will be selected. (e.g. for an interval [170, -170] -> both [170, 180[ and [-180, -170] intervals will be used to list/subset data).
To list files corresponding to half orbits crossing the bounding box:
bbox = -126, 32, -120, 40
fc.list_files(
version='PIC?',
subset='Basic',
bbox=bbox)
| cycle_number | pass_number | time | subset | version | filename | |
|---|---|---|---|---|---|---|
| 0 | 9 | 11 | [2024-01-04T12:08:54.000000, 2024-01-04T12:59:... | ProductSubset.Basic | PIC0_01 | /home/runner/work/fcollections/fcollections/do... |
| 1 | 11 | 11 | [2024-02-15T05:39:04.000000, 2024-02-15T06:29:... | ProductSubset.Basic | PIC0_01 | /home/runner/work/fcollections/fcollections/do... |
| 2 | 10 | 11 | [2024-01-25T08:54:00.000000, 2024-01-25T09:44:... | ProductSubset.Basic | PIC0_01 | /home/runner/work/fcollections/fcollections/do... |
To query a subset of Swot LR L2 data crossing the bounding box:
Note
Lines of the swath crossing the bounding box will be entirely selected.
bbox = -126, 32, -120, 40
ds_area = fc.query(subset="Basic", version='P?C?', cycle_number=9, pass_number=11, bbox=bbox)
# Figure
localbox_cartopy = bbox[0] - 1, bbox[2] + 1, bbox[1] - 1, bbox[3] + 1
fig, ax = plt.subplots(figsize=(8, 8), subplot_kw=dict(projection=ccrs.PlateCarree()))
ax.set_extent(localbox_cartopy)
plot_kwargs = dict(
x="longitude",
y="latitude",
cmap="Spectral_r",
vmin=-1.5,
vmax=1.5,
cbar_kwargs={"shrink": 0.3},)
# SWOT KaRIn SLA plots
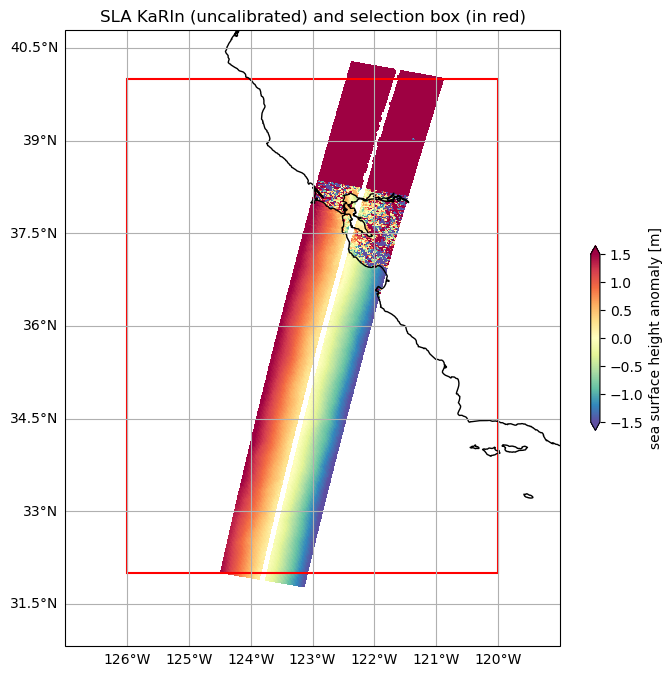
ds_area.ssha_karin_2.plot.pcolormesh(ax=ax, **plot_kwargs)
ax.set_title("SLA KaRIn (uncalibrated) and selection box (in red)")
ax.coastlines()
ax.gridlines(draw_labels=['left', 'bottom'])
# Add the patch to the Axes
rect = patches.Rectangle((bbox[0], bbox[1]), bbox[2] - bbox[0], bbox[3] - bbox[1], linewidth=1.5, edgecolor='r', facecolor='none')
ax.add_patch(rect)
<matplotlib.patches.Rectangle at 0x7f36f43bfe00>
Swath sides in Level-2 Unsmoothed subset#
The L2_LR_SSH Unsmoothed dataset files are using netcdf groups to separate the
swath sides. This means we can open one of the two sides. The following figure
illustrates how the left_swath and right_swath parameters can be used to
retrieve one or the other side.
ds_left = fc.query(subset='Unsmoothed', cycle_number=9, pass_number=10, left_swath=True, right_swath=False,
selected_variables=['longitude', 'latitude', 'sig0_karin_2']).compute()
ds_right = fc.query(subset='Unsmoothed', cycle_number=9, pass_number=10, left_swath=False, right_swath=True,
selected_variables=['longitude', 'latitude', 'sig0_karin_2']).compute()
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 4))
plot_kwargs = dict(
cmap="Greys_r",
cbar_kwargs={"shrink": 0.3},
vmin=5, vmax=60)
# SWOT KaRIn SLA plots
s = slice(45000, 50000, 3)
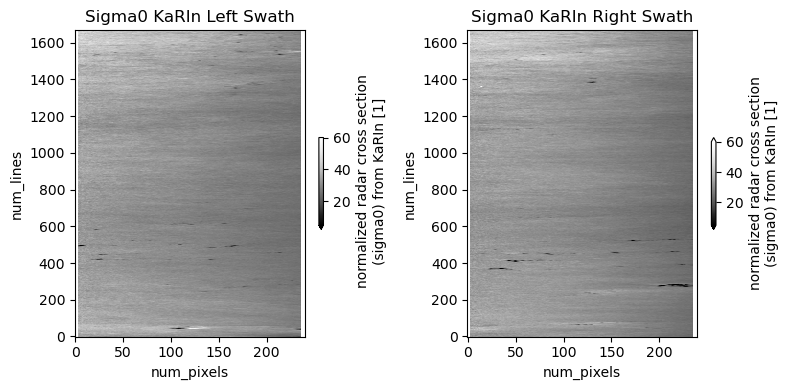
ds_left.isel(num_lines=s).sig0_karin_2.plot.imshow(ax=ax1, **plot_kwargs)
ds_right.isel(num_lines=s).sig0_karin_2.plot.imshow(ax=ax2, **plot_kwargs)
ax1.set_title("Sigma0 KaRIn Left Swath")
ax2.set_title("Sigma0 KaRIn Right Swath")
fig.tight_layout()
Note
Combination of both sides is not yet possible. Moreover, keep in mind that position coordinates may include invalids which can break geo-plots.
